Kroksveen AC, Jaffe JD, Aasebø E, Barsnes H, Bjørlykke Y, Franciotta D, Keshishian H, Myhr KM, Opsahl JA, van Pesch V, Teunissen CE, Torkildsen Ø, Ulvik RJ, Vethe H, Carr SA, Berven FS.
Proteomics. 2015. doi: 10.1002/pmic.201400142. [Epub ahead of print]
Multiple sclerosis (MS) is a chronic inflammatory disease of the CNS with unknown cause. Proteins with different abundance in the cerebrospinal fluid (CSF) from relapsing-remitting MS (RRMS) patients and neurological controls could give novel insight to the MS pathogenesis and be used to improve diagnosis, predict prognosis and disease course, and guide in therapy decisions. We combined iTRAQ labeling and Orbitrap mass spectrometry to discover proteins with different CSF abundance between six RRMS patients and 18 neurological disease controls. From 777 quantified proteins seven were selected as biomarker candidates, namely chitinase-3-like protein 1, secretogranin-1 (Sg1), cerebellin-1, neuroserpin, cell surface glycoprotein MUC18, testican-2, and glutamate receptor 4. An independent sample set of 13 early-MS patients, 13 RRMS patients and 13 neurological controls was used in a multiple reaction monitoring verification study. We found the intracellular calcium binding protein Sg1 to be increased in early-MS patients compared to RRMS and neurological controls. Sg1 should be included in further studies to elucidate its role in the early phases of MS pathogenesis and its potential as a biomarker for this disease.
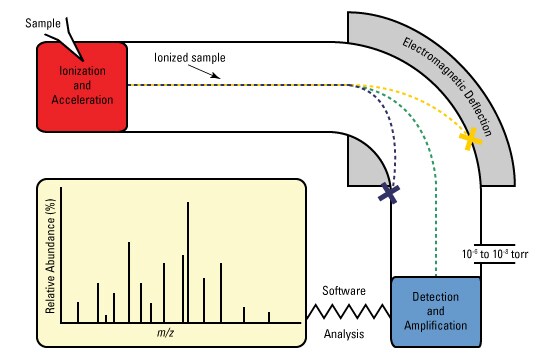
Figure: The mass spectrometer. The sample is loaded into one end and the molecules which make up the sample are then turned into ions (the process of ionization). The ions are electrically charged and are deflected by the magnetic field. The relative speeds at which they hit the detector at the end then depends on their relative charge and mass (how heavy they are). A computer then converts them to a readable format where peptides can be identified.
Secretogranin 1, also known as Chromogranin B is a neuroendocrine secretory granule protein and like other granins contributes to the formation of the secretory vesicle. All proteins are secreted from cells, the rate of synthesis of the secreted substance. It is not readily apparent what the relevance of this protein in MS is, however, the authors believe that secretogranin may prove to be a good predictor of MS disease progression in the future, but more work is needed here. Conversely, in schizophrenia low levels of this protein has been detected, but here there is a genetic linkage study to prove its relevance in schizophrenia.
One of the other proteins they've detected is chitinase-3-like protein 1; there is an increasing body of evidence that this can predict conversion from CIS (clinically isolated syndrome, a single demyelinatory event) to MS.
I have one major problem with this work, that is that the authors have confirmed their finding by doing a fresh set of MS samples again using mass spectrometry, i.e. a similar methodology. In biomarker verification it is important to reproduce the findings using two different methodologies. Most often this is developing your own enzyme-linked immunoassay, which is the most commonly used methodology in verification studies. Now the job of proving or disproving the findings in a larger sample set becomes someone else's job!
However, this work does raise the issue that we know/understand less about the biological disease process in MS than we should. This sentence in its various reincarnations seems to be a repeated mantra from me in this blog.
Labels: chromogranin B, proteomics, secretogranin-1
